Resources
Internally Developed Tools

software
Pangolin
Pangolin was developed to implement the dynamic nomenclature of SARS-CoV-2 lineages, known as the Pango nomenclature. It allows a user to assign a SARS-CoV-2 genome sequence the most likely lineage (Pango lineage) to SARS-CoV-2 query sequences.
Pangolin assigns lineages to query sequences as described in Rambaut et al 2020.
View

software
Scorpio
Serious constellations of reoccurring phylogenetically-independent origin. A tool for snp-based calling of variants of concern.
View

software
Civet
Civet is a tool developed with 'real-time' genomics in mind.
Using a background phylogeny, such as the large phylogeny available through the COG-UK infrastructure on CLIMB, civet will generate a report for a set of sequences of interest i.e. an outbreak investigation.
View

software
Polecat
Using a background phylogeny, such as the large phylogeny available through the COG-UK infrastructure on CLIMB, polecat will identify and flag clusters based on various configurable statistics.
View

website
pango.network
A website documenting the Pango nomenclature as well as the policies involved with designating new lineages.
View
Dashboards

Su, Wu, and Andersen labs at Scripps Research
Outbreak.info
A data dashboard for viewing more information about the various SARS-COV-2 variants
View

Emma Hodcroft, University of Bern, Switzerland
CoVariants
A dashboard giving an overview of SARS-CoV-2 variants and interesting mutations
View

Computational Evolution group at ETH Zurich, Switzerland
covSPECTRUM
An interactive dashboard SARS-CoV-2 variants with sophisticated queries
View

Center for Systems Science and Engineering at Johns Hopkins University
JHU Covid-19 Dashboard
An interactive dashboard displaying various information about the general spread of Covid-19.
View
Kieran Lamb @ CVR Glasgow
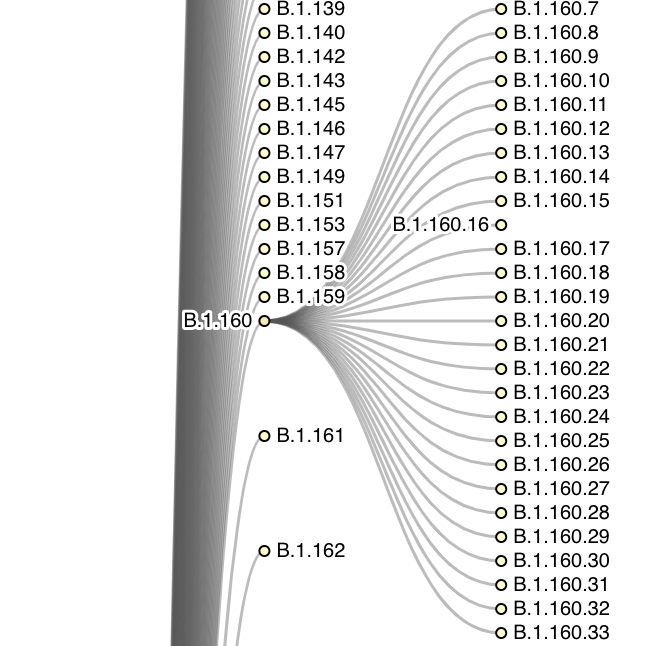
Lineage Tree
An interactive visualisation representing the Pango lineage system through a collapsable tree structure.
View

COG-UK
COG Mutation Explorer
A data visualisation tool for exploring important SARS-CoV-2 variants and mutations
View
Software Tools

Zuher Jahshan, Leonid Yavits
CoViT
A tool that enables real-time phylogenetics for the SARS-CoV-2 pandemic using Vision Transformers
View

Centre for Genomic Pathogen Surveillance
PanGUIlin
A web application version of the Pangolin Tool
View

Trevor Bedford et al
Nextstrain
Powerful, interactive tools for exploring virus data. Nextstrain provides a large database of viral genomes and bioinformatics pipelines for phylogenetic analysis, including interactive tree visualisations.
View

Angie Hinrichs, UC Santa Cruz Genomics Institute
UShER Phylogenetic Placement
Upload your SARS-CoV-2 sequence (FASTA or VCF file) to find the most similar complete, high-coverage samples from GISAID or from public sequence databases (NCBI Virus / GenBank, COG-UK and the China National Center for Bioinformation), and your sequence's placement in the UCSC/UShER phylogenetic tree.
View
Data Sources

GISAID
GISAID
Important, global data repository for SARS-COV-2 genomes as well as for other pathogens
View

ENA
European Nucleotide Archive
The ENA provides a high quality, downloadable data set of SARS-CoV-2 genomes. ENA data is available as a data source on Galaxy.
View

COG-UK
COG-UK
COG-UK provides a large number of data sets and bioinformatics tools for SARS-COV-2.
View